The new 2026 rectal cancer guidelines now include organ preservation and suggest that radiologists will see increased follow-up MRI, according to a March 6 multidisciplinary education session at ECR 2026.
The guidelines, published on 29 January, focus more on risk-adapted staging and restaging driven by treatment stratification, said Dr. Regina Beets-Tan, PhD, from Maastricht University, the Netherlands. For radiologists, evaluating tumor response will include multiple time points, she added.
"Whether you want it or not, the rectal cancer radiologist should know this field of organ preservation as it is included in the international clinical practice guidelines," Beets-Tan stressed.
 Dr. Regina Beets-Tan.The ESR.
Dr. Regina Beets-Tan.The ESR.
Organ preservation using new neoadjuvant treatment strategies is seen as an alternative to surgery for some patients, but the best timing of the decision, based on treatment response, is not well understood, according to Beets-Tan. The guidelines prompt delving much deeper into the question of clinical response -- whether it is complete or whether it is a good response but not clinically complete.
Diffusion-weighted imaging MRI has been shown to be the most effective and most accurate method for identifying complete responders in a fibrotically changed irradiated tumor bed, Beets-Tan explained during the talk.
"But if we look at whether diffusion MRI is better than endoscopy, then the single most accurate method is endoscopy," she said. "If we combine all modalities, then we can select the patient the most accurate way. It's about a multidisciplinary approach, and that is reflected in the guidelines.
"Sometimes we have the endoscopy at our disposal, sometimes we need to talk with the team about what was found at endoscopy," Beets-Tan noted. "It is the entire multidisciplinary team that needs to discuss and stratify the right treatment."
Imaging criteria will also need to evolve as more rectal cancer treatments become available -- immunotherapies, combination therapies of immunotherapy with radiotherapy, and radiotherapy with chemotherapy, or chemotherapy as a standalone treatment, Beets-Tan added.
"All these treatments are in the trial setting at the moment, but we already know that the imaging criteria are different," she said.
Ultimately, "we have to focus our research on looking at other biomarkers, clinical, maybe blood, maybe tissue, genetics, but always combine it with imaging," she said. In the end, the medical community may see predictive outcome models.
The session featured the radiation oncologist perspective of Dr. Vincenzo Valentini from Gemelli ART (Advanced Radiation Therapy) and the Catholic University of the Sacred Heart in Rome, Italy, and the surgical perspective of Dr. Geerard Beets, also from Maastricht.
The guideline document was published in European Radiology.
Our full coverage of ECR 2026 can be found here.




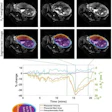
![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnieeurope.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=100&q=70&w=100)





![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnieeurope.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=112&q=70&w=112)