Quantification of the well-aerated lung at baseline chest CT can be a valuable tool for predicting prognosis in patients with COVID-19 pneumonia, a team of Italian researchers reported in a study published online on 17 April in Radiology.
In the study involving over 200 patients, quantification of well-aerated lung parenchyma -- either visually by radiologists or via software -- on baseline chest CT was found to predict which patients would be admitted to the intensive care unit (ICU) or death in patients with COVID-19 pneumonia.
"Quantitative assessment of the extent of lung involvement by COVID‑19 pneumonia may be useful for routine patient management," concluded the researchers led by Dr. Davide Colombi of Guglielmo da Saliceto Hospital in Piacenza, Italy.
Seeking to assess the prognostic value of these measurements, the researchers retrospectively analyzed patients who were admitted to their institution's emergency department between 17 February and 10 March with suspicion of COVID-19 pneumonia. The researchers first quantified well-aerated lung parenchyma on baseline CT studies for the remaining 236 patients visually as an estimate of the percentage of well-aerated lung. The open-source 3D-Slicer software's Chest Imaging Platform extension was then used to calculate that same measurement, as well as the absolute volume of well-aerated lung parenchyma and the volume of adipose tissue area.
Next, the researchers performed logistic regression analysis to assess the relationship of these CT metrics and clinical parameters (demographics, comorbidities, symptoms and symptom duration, oxygen saturation, and laboratory values) with patient outcomes. Of the patients who were admitted to the ICU or died, 16% had four or more affected lung lobes at baseline CT, compared with 6% who didn't. That difference was statistically significant (p < 0.04).
They also found that a visual assessment of less than 73% well‑aerated lung parenchyma (odds ratio: 5.4; p < 0.001), a software assessment of less than 71% well-aerated lung parenchyma (odds ratio: 3.8; p < 0.001), and a software assessment of less than 2.9 L of well-aerated parenchyma total volume (odds ratio: 2.6; p < 0.01) were all predictors of ICU admission or death. The authors then calculated performance for three different quantitative models based on these parameters.
| Performance for predicting ICU admission/death | ||||
| Model with clinical parameters only | Model with clinical parameters and a visual assessment of < 73% well-aerated lung parenchyma | Model with clinical parameters, software-based assessment of < 71% well-aerated lung parenchyma, and > 262 cm2 of adipose tissue area volume | Model with clinical parameters, software-based assessment of < 2.9 L of absolute volume of well-aerated lung parenchyma, and > 262 cm2 of adipose tissue area volume | |
| Sensitivity | 75% | 72% | 75% | 75% |
| Specificity | 73% | 81% | 80% | 81% |
| Positive predictive value | 70% | 76% | 75% | 77% |
| Negative predictive value | 78% | 78% | 80% | 79% |
| Area under the curve | 0.83 | 0.86 | 0.86 | 0.86 |
The improved performance over the clinical parameters-only model was statistically significant (p < 0.04) for both the model with visual assessment and the model that included software-based assessment of lung parenchyma absolute volume.
"Visual and software-based quantification of well-aerated lung parenchyma on admission chest CT were predictors of [ICU] admission or death in patients with COVID-19 pneumonia," the authors concluded.


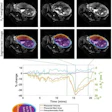
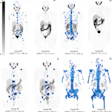
![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnieeurope.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=100&q=70&w=100)






![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnieeurope.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=112&q=70&w=112)