Radiomics scoring derived from contrast-enhanced MRI can predict pathologic complete response in women being treated for triple-negative breast cancer, according to findings published on 3 September in Radiology.
Researchers led by Dr. Toulsie Ramtohul, from the Curie Institute in Paris found that a radiomics score at pretreatment perfusion MRI performed well in predicting pathologic complete response in these patients, who were being treated with immunotherapy-based regimens. They highlighted that this method has the potential to serve as a biomarker for drug sensitivity heterogeneity in studies that explore less aggressive breast cancer treatment.
“The proposed radiomics score, derived from images obtained using various ‘real-life’ MRI scanner models, demonstrated favorable discrimination and clinical utility for predicting pathologic complete response,” Ramtohul and colleagues wrote.
Radiology researchers over the years have attempted to develop effective methods for predicting how patients being treated for breast cancer may respond to treatment strategies. The researchers noted that predictors of response to neoadjuvant chemotherapy have not been identified for women being treated for triple-negative breast cancer.
Ramtohul and co-authors studied how radiomics derived from pretreatment perfusion MRI could serve as a predictive marker for pathologic complete response in these patients.
The team obtained pretreatment dynamic contrast-enhanced MRI (DCE-MRI) scans by using scanners from multiple vendors. It also used the Tofts model to segment tumors and analyze pharmacokinetic parameters. The following radiomics features were extracted from the images: rate constant for contrast agent plasma-to-interstitial transfer (or Ktrans), volume fraction of extravascular and extracellular space (Ve), and maximum contrast agent uptake rate (Slopemax) maps.
The study included 195 women with triple-negative breast cancer who underwent neoadjuvant chemotherapy and were enrolled between 2021 and 2023. The researchers assessed their radiomics model’s performance in a training set (n = 112) and a test set (n = 83).
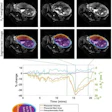
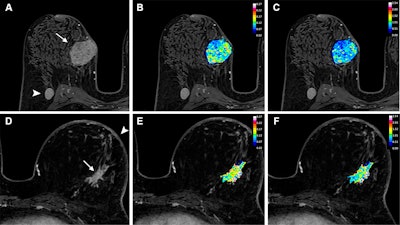 Axial dynamic contrast-enhanced MRI scans show results from two women before they underwent neoadjuvant chemoimmunotherapy for triple-negative breast cancer. (A) T1-enhanced MRI scan, (B) rate constant for contrast agent plasma-to-interstitial transfer (Ktrans) map, and (C) maximum contrast agent uptake rate (Slopemax) map in a 56-year-old woman depict a tumor that manifested as a spherical 4-cm mass enhancement (arrow in A) in the right breast with axillary node involvement (arrowhead in A). The radiomics score was high, at 3.3. Histopathologic examination after chemotherapy revealed no invasive residual cancer in the breast or the lymph nodes (pathologic complete response). (D) T1-enhanced MRI scan, (E) Ktrans map, and (F) Slopemax map in a 49-year-old woman show a tumor that manifested as a spiculated 3-cm mass enhancement (arrow in D) in the left breast with thickening of the skin (arrowhead in D). The parametric maps show much greater spatial heterogeneity than in the case shown in B and C, with areas of high intensity. The radiomics score was low, at −1.6. Histopathologic examination after chemotherapy revealed a 10-mm invasive residual cancer in the breast (not pathologic complete response). Image courtesy of the RSNA.
Axial dynamic contrast-enhanced MRI scans show results from two women before they underwent neoadjuvant chemoimmunotherapy for triple-negative breast cancer. (A) T1-enhanced MRI scan, (B) rate constant for contrast agent plasma-to-interstitial transfer (Ktrans) map, and (C) maximum contrast agent uptake rate (Slopemax) map in a 56-year-old woman depict a tumor that manifested as a spherical 4-cm mass enhancement (arrow in A) in the right breast with axillary node involvement (arrowhead in A). The radiomics score was high, at 3.3. Histopathologic examination after chemotherapy revealed no invasive residual cancer in the breast or the lymph nodes (pathologic complete response). (D) T1-enhanced MRI scan, (E) Ktrans map, and (F) Slopemax map in a 49-year-old woman show a tumor that manifested as a spiculated 3-cm mass enhancement (arrow in D) in the left breast with thickening of the skin (arrowhead in D). The parametric maps show much greater spatial heterogeneity than in the case shown in B and C, with areas of high intensity. The radiomics score was low, at −1.6. Histopathologic examination after chemotherapy revealed a 10-mm invasive residual cancer in the breast (not pathologic complete response). Image courtesy of the RSNA.
The radiomics score achieved area under the curve (AUC) values of 0.84 for the training set and 0.8 for the test set in predicting pathologic complete response. A nomogram incorporating the radiomics score, grade, and Ki-67, meanwhile, achieved an AUC of 0.86 in the test set.
The team also observed significant associations between higher radiomics scores (< 0.25) and the following characteristics: tumor size (p < 0.001), washout enhancement (p = 0.01), androgen receptor expression (p = 0.009), and programmed death ligand 1 expression (p = 0.01). It highlighted that this shows a correlation with tumor immune environment in participants with triple-negative breast cancer.
The study authors highlighted that their radiomics score could potentially be used to improve clinical decision-making.
In an accompanying editorial, Dr. Gaiane Rauch, PhD, from the University of Texas MD Anderson Cancer Center in Houston wrote that the results by Ramtohul et al “confirm the importance of including tumor microenvironment biomarkers” in image-based models. This can further guide escalation or deescalation of neoadjuvant therapy in patients with triple-negative breast cancer.”
“Large-scale multicenter clinical trials that include functional imaging, clinicopathologic, and genomic data are needed,” Rauch added. “The long-term goal is to develop a personalized, response-adapted, biomarker-driven approach to optimize neoadjuvant therapy and improve quality of life and outcomes in patients with triple-negative breast cancer.”
The full results can be found here.


![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnieeurope.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=100&q=70&w=100)




![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnieeurope.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=112&q=70&w=112)