Using an AI tool with brain MRI in children finds 64% of focal cortical dysplasias (FCDs) -- abnormalities linked to epilepsy -- that radiologists miss, researchers have reported.
The tool, called MELD Graph, improves detection of FCDs, which are a leading cause of epilepsy, according to a team led by Mathilde Ripart, PhD, of the University College London in the U.K. Using it with brain MRI could speed up time to diagnosis and allow patients to get the surgical treatment they need more quickly while also reducing costs to the healthcare system by almost $70,000 (€67,000) per patient, the group noted. The study was published on 24 February in JAMA Neurology.
"[MELD Graph's] clinical implementation holds promise for early diagnosis and improved management of focal epilepsy, potentially leading to better patient outcomes," Ripart and colleagues wrote.
FCDs are a common structural cause of epilepsy and are difficult to control with medication; however, once identified, they can be treated surgically. But they can be difficult to see and up to half are missed by radiologists' brain MR imaging -- which can translate to delays in diagnosis and surgery, more seizures and visits to the emergency room, and disruption of school and home life, they explained.
The group investigated whether an AI tool could help detect these lesions on MRI via a study that included data from 1,185 participants (703 people with FCD and 482 controls) from 23 epilepsy centers around the world. The data came from the Multicenter Epilepsy Lesion Detection project (MELD, conducted between 2018 and 2022), and half of the participants were children. Data from 20 of these centers were split equally between training and testing cohorts, and data from three centers were used for "site-independent" testing.
The tool is called MELD Graph, and is a "context-aware graph" neural network based on nnU-Net, the researchers noted. It segments FCDs by integrating information "across whole cortical hemispheres," and in this study collated 34 surface-based MRI features and manual lesion masks (the latter being outlines of lesions in brain images that are usually created by hand) from study participants. The investigators compared the performance of MELD Graph for identifying FCDs to another AI algorithm they previously created that is a patch-based (that is, 1-cm3 patches of cortex) multilayer perceptron called MELD MLP.
They reported the following:
| Comparison of performance between MELD MLP to MELD Graph for identifying FCDs | ||
|---|---|---|
| Test dataset | MELD MLP | MELD Graph |
| Sensitivity | 67% | 70% |
| Specificity | 54% | 60% |
| Positive predictive value | 39% | 67% |
The group also found that, in the test dataset, MELD Graph had a sensitivity of 81.6% in patients who were seizure-free a year after surgery and 63.7% in MRI-negative patients with FCD.
Although MELD Graph is not yet clinically available, Ripart and colleagues have released it as open-source software, and are training clinicians and researchers around the world on its use.
"One of the highlights for me [of developing MELD Graph] is hearing from doctors around the world -- including the U.K., Chile, India, and France -- who have been able to use [it] to help their own patients," Ripart said.
The complete study can be found here.



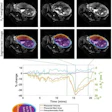
![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnieeurope.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=100&q=70&w=100)




![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnieeurope.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=112&q=70&w=112)