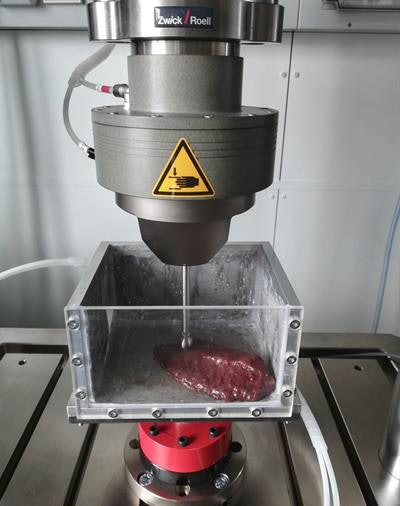
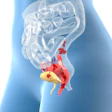
Researchers from Austria have set out to create 3D-printed organs using materials that resemble the properties of real tissue.
In recent years, clinicians have been using 3D-printed anatomical models to plan and simulate complex surgeries. One of the main drawbacks of this practice is that the models lack the realistic characteristics of biological tissue, noted lead investigator Dieter Pahr, PhD, from Karl Landsteiner University of Health Sciences in Krems.
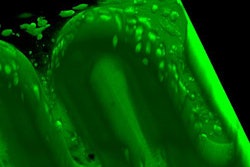
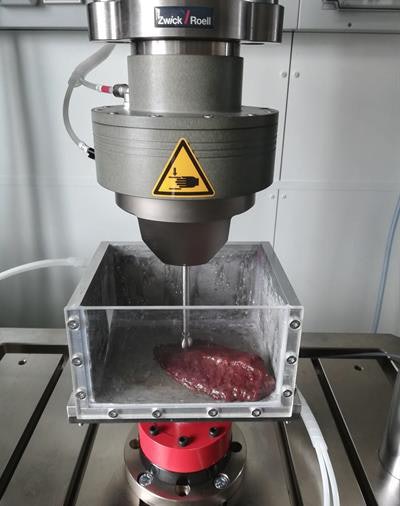 Identifying the biomechanical properties of organ tissue for 3D bioprinting. Image courtesy of Dieter Pahr, PhD, from Karl Landsteiner University of Health Sciences in Krems.
Identifying the biomechanical properties of organ tissue for 3D bioprinting. Image courtesy of Dieter Pahr, PhD, from Karl Landsteiner University of Health Sciences in Krems.Hoping to improve the value of 3D-printed models, Pahr and colleagues have begun a new research project at their institution that aims to identify the biomechanical properties of actual organ tissue and then use this information to determine the best-suited biomaterials for making lifelike 3D-printed organs.
Furthermore, they plan to develop a computer model that can predict the mechanical qualities of 3D-printed biological tissues based on their microstructures and their material composition. Ultimately, the group hopes that by inputting the properties of a particular type of tissue into a computer program, they will be able to create realistic 3D-printed organs.
This work can bolster the field of medical 3D printing and may eventually "open the door for highly customized and realistic tissue and organ models for medical use," the researchers noted.








![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnieeurope.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=112&q=70&w=112)