The use of thin-client 3D technology can facilitate offsite reading of CT angiography (CTA) studies in on-call situations, according to research from Charité - Campus Benjamin Franklin in Berlin.
"With a thin-client server design, typical bottlenecks such as data transfer, computing power, and user memory of a workstation solution have been overcome," said Dr. Bernhard Meyer. "Therefore, sophisticated postprocessing can be performed from home on a basic laptop or desktop computer."
Meyer presented the research team's findings during a scientific session at the European Congress of Radiology (ECR) in Vienna.
To evaluate the benefits of thin-client technology for offsite expert reading of runoff CTA exams in an on-call environment, the researchers studied 20 CTA exams using Visage CS thin-client advanced visualization software (Visage Imaging, Andover, MA) and a DSL with 6 Mb/sec of bandwidth.
To assess the feasibility of system use by the interventional radiologist on-call at home, the researchers recorded the time from the initial phone call to study availability on the software client. The study team also measured the time to final diagnosis and compared the results to the processing time for studies read in the hospital.
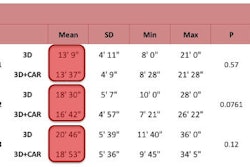
With the offsite method, users could interact with diagnostic-quality images in less than one minute, Meyer said. The time from the initial phone call to the interventional radiologist receiving the study on the screen ranged from 2.5 to 11 minutes, with a median of six minutes. Postprocessing time ranged from 5.5 to 16 minutes, with a median of 12.5 minutes.
The time from the initial phone call to when the final diagnosis was given to the resident ranged from 12.5 to 21 minutes, with a median of 15 minutes. No significant time difference was noted for offsite postprocessing and in-hospital postprocessing (p > 0.05), Meyer said.
In addition, there was 100% agreement between onsite and offsite reading in the therapeutic measures taken, Meyer said.
"This thin-client solution provides high reliability and performance comparable to the onsite setting, and can be routinely used for image postprocessing from home in an on-call situation," he said.
By Erik L. Ridley
AuntMinnie.com staff writer
April 23, 2009
Related Reading
3D plus CT improves imaging of asbestos-related diseases, April 16, 2009
Computer pen bests mouse for 3D image segmentation, February 20, 2009
Hybrid 3D echo/SPECT technique produces better CAD imaging, February 12, 2009
Virtual colonoscopy beginners do better with 3D data, February 5, 2009
3D image guidance improves spinal surgery outcomes, December 29, 2008
Copyright © 2009 AuntMinnie.com








![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnieeurope.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=112&q=70&w=112)