A potential new weapon in the arsenal for treating prostate cancer is being discussed at this week's SNM annual meeting in Toronto.
Alpha particle-emitting radiopeptides have proved effective in treating prostate cancer in mice, tripling their average survival rate, researchers reported. Lead author Dr. Damian Wild of the University Hospital Basel in Switzerland stated that this novel form of treatment has the potential to target and destroy cancer cells with minimum damage to surrounding healthy tissue.
Researchers have been able to develop highly specific radiopeptides that bind with tumor cells and treat the tumors with specific therapeutic radioactive substances attached to the radiopeptide. Prostate cancer cells have an overabundance of gastrin-related peptide receptors, which makes this cancer a strong candidate for treatment.
The study conducted by the Swiss researchers compared the outcomes of two different types of radiopeptides with a control group that received no treatment at all. Mice receiving the 213 Bi-DOTA PESIN peptide at the maximum tolerated dose had the highest survival rate.
More than 186,000 men in the U.S. are diagnosed with prostate cancer each year, and more than 30,000 previously diagnosed patients experience cancer recurrence. The cancer in most recurrences cannot be localized and treated adequately with conventional treatments. The researchers hope that their discovery will lead to a systemic treatment that efficiently kills small tumors.
Related Reading
Longer hormone therapy may improve outcomes in advanced prostate cancer, April 15, 2009
SBRT for early prostate cancer shows promising interim results, April 9, 2009
Adding radiotherapy to hormone treatment cuts prostate cancer mortality, December 16, 2008
New tracer shows promise for PET/CT prostate imaging, January 18, 2007
Copyright © 2009 AuntMinnie.com


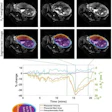
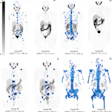
![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnieeurope.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=100&q=70&w=100)





![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnieeurope.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=112&q=70&w=112)