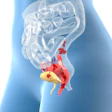
The first large-scale study of whole-genome testing identified those breast cancer patients who might benefit the most from certain treatments, according to new research published online in Lancet Oncology.
The researchers from several institutions in France tested all the DNA of tumor cells from more than 400 women with advanced breast cancer. The goal was to see whether whole-genome analysis could identify unique characteristics and abnormal genes in the metastatic tissue, which could then be targeted for treatment in later trials.
Dr. Fabrice André, from Institut Gustave Roussy, and colleagues looked at the quality of tumor samples, the proportion of women for whom the genomic analyses could be done, and the proportion for whom a targeted therapy could be offered. They took biopsy samples from 407 patients from 18 centers across France (Lancet Oncology, February 7, 2014).
The group was able to perform whole-genome analysis in two-thirds of the patients. Forty-six percent of the 423 enrolled patients were found to have a targetable genomic alteration, while 39% had a rare alteration.
To date, 28% of the women with targetable alterations have been matched with new treatments that are being tested in clinical trials, André said in a statement released by the journal. The team's goal is to have 30% of the enrolled patients in clinical trials testing therapies targeting the alterations identified in their tumors.
"Our findings indicate that large molecular screening programs, performed in the context of clinical trials, are helpful to see whether a patient with metastatic cancer could be eligible for a targeted therapy matched to a genomic alteration," André said.











![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnieeurope.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=112&q=70&w=112)