Providing readers with individual feedback on their performance after using chest radiography computer-aided detection (CAD) software does not yield an improved ability to differentiate CAD candidates for pulmonary nodules, Dutch researchers have found.
Testing their hypothesis, that providing short-term feedback to readers on their performance would improve their confidence in CAD analysis, and their ability to differentiate between true-positive and false-positive CAD candidates, the Academic Medical Center (AMC) team found that such feedback failed to yield the hoped-for statistical improvements.
"Short-term feedback does not increase the ability of readers to differentiate true- from false-positive candidate lesions and to use CAD more effectively," wrote a research team led by Dr. Diederick De Boo of the radiology department from AMC in the Netherlands.
Previous research from AMC had found that readers using CAD for pulmonary nodules in chest radiography as a second reader had difficulties in distinguishing between true-positive and false-positive CAD candidates, De Boo said.
"False-positive candidates were accepted and true-positive candidates were dismissed," De Boo told AuntMinnieEurope.com. "This was especially true for subtle to very subtle nodules/cancers."
In their current retrospective study published in European Radiology, the AMC team explored the potential for individual feedback to improve reader effectiveness with CAD. They selected 140 patients from their institution's archive, of which 56 had a solitary CT-proven nodule and 84 were negative controls (Eur Radiol, June 2012, Vol. 22:8, pp. 1659-1664).
The study included patients who had a two-view chest radiograph and thoracic CT within six weeks and had no opacity or a single nodular opacity. Each of the chest x-rays were divided into four subsets of 35, which were each read by six readers. The six readers in the study included five radiology residents with zero to five years of training, and one board-certified radiologist with more than 15 years of experience interpreting chest films, according to the authors.
Each subset, which included 14 cases with a single nodular opacity and 21 negative control cases, were interpreted first with and then without the use of CAD (IQQA-Chest, EDDA Technology) as a second reader. They were asked to determine the presence or absence of an intrapulmonary opacity using a five-point scale of confidence ranging from five (pulmonary nodule definitely present) to three (equivocal with respect to the presence of a nodule) and one (definitely no nodule present).
If a confidence rating of greater than one was given, observers were asked to indicate the location of the suspected lesion on a data sheet. All readers were informed that images contained only a single nodular opacity and were told to ignore calcified lesions and lesions smaller than 5 mm in diameter. Readers were allowed to magnify, adjust window/level, and reverse grayscale levels during both readings.
After each subset, the readers received individual feedback from a researcher. On a case-by-case basis, the researcher and reader discussed the location of the lesions, CAD marks if present with respect to whether they were true-positive or false-positive, and the reader's individual response, according to the study team.
The researchers then calculated sensitivity, specificity, and area under the receiver-operating characteristics curve (AUC) for interpretations with and without CAD. The standalone CAD sensitivity was 59%, with 1.9 false-positives per image.
While the mean reader AUC and sensitivity increased slightly after using CAD, the differences were not statistically significant. Specificity dropped slightly after access to CAD, but this result was also not significant.
Mean reader performance, with and without CAD
|
All six readers dismissed 32 (17%) of the 192 true-positive CAD candidates. No difference was seen between the first two subsets read and the last two. While the number of false-positive CAD marks that were accepted by readers declined from 10 in the first two subsets to six in the last two, that difference was not statistically significant.
"It should be mentioned that the [study] results only apply for the current tested CAD systems, and systems are improved as we speak," De Boo noted. "So the results may not [apply] to future systems or other reader groups/patients groups."
In addition, the authors said "further research is needed to determine whether a longer training period or additional processing and display tools such as temporal subtraction, rib suppression, or likelihood calculations can increase the benefits of CAD for reader performance in chest radiography."







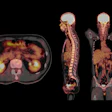

![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnieeurope.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=112&q=70&w=112)