WASHINGTON (Reuters), Oct 9 - British researchers said they have found a new genetic mutation that doubled the risk of breast cancer in women who carry it.
The gene, called BRIP1, helps to repair damaged DNA -- like some of the other known breast cancer genes, researchers reported in this week's issue of the journal Nature Genetics.
And, as with the BRCA2 breast cancer gene, certain mutations in BRIP1 may cause a blood disease called Fanconi anemia, reported Dr. Nazneen Rahman of the Institute of Cancer Research, in Sutton, Britain and colleagues.
"Breast cancer is approximately twice as common in sisters and mothers of affected individuals as in the general population," the researchers wrote.
Mutations in BRCA1, BRCA2, and TP53 raise the risk of breast cancer by 10- to 20-fold by age 60. Mutations in genes called CHEK2 and ATM roughly double the risk.
"Together, these susceptibility genes are estimated to account for up to 25% of the familial risk of breast cancer," the researchers wrote.
They looked for genes that interact with the known breast cancer genes and found BRIP1, also known as BACH1.
They screened 1,212 women with breast cancer who did not have mutations in BRCA1 and BRCA2 and more than 2,000 women without breast cancer.
Nine of the breast cancer patients had mutations in the BRIP1 gene but just two of 2,081 cancer-free women did, Rahman's team found.
Breast cancer kills 500,000 people a year globally according to the World Health Organization.
Last Updated: 2006-10-09 12:01:09 -0400 (Reuters Health)
Related Reading
Copyright © 2006 Reuters Limited. All rights reserved. Republication or redistribution of Reuters content, including by framing or similar means, is expressly prohibited without the prior written consent of Reuters. Reuters shall not be liable for any errors or delays in the content, or for any actions taken in reliance thereon. Reuters and the Reuters sphere logo are registered trademarks and trademarks of the Reuters group of companies around the world.






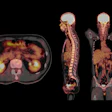

![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnieeurope.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=112&q=70&w=112)