Portuguese researchers have used SPECT to detect Parkinson's disease with 98% accuracy by employing quantitative, automated analysis based on machine learning. Validated using brain images from 654 individuals, the approach outperformed visual analysis and other automated techniques in the literature, and can enable more appropriate and timely treatments for patients.
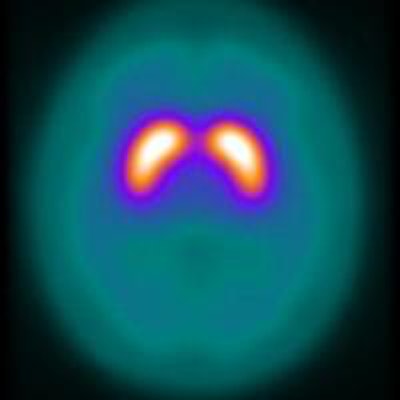
Dr. Miguel Castelo-Branco, PhD, and Francisco Oliveira, PhD, of the University of Coimbra applied their technique to iodine-123 labeled FP-CIT SPECT, a popular tool in the diagnosis of Parkinson's disease. In the scans, decreased uptake of the cocaine analogue in the brain's striatum indicates an absence of the dopamine transporters that the tracer would otherwise bind to in disease-free individuals. Images are used to exclude conditions including essential tremor. However, in some cases, especially those with early stage Parkinson's, changes in uptake can be difficult to detect by eye.
 Miguel Castelo-Branco and colleagues at the University of Coimbra's Institute for Nuclear Sciences Applied to Health.
Miguel Castelo-Branco and colleagues at the University of Coimbra's Institute for Nuclear Sciences Applied to Health.Algorithm learns from images
The new approach uses support vector machines, a popular machine learning technique. It uses an algorithm trained with a set of images of known status, where each is characterized by a set of preselected quantitative features. Each image voxel contained in striatal volumes-of-interest (VOI) on the left and right side of the brain constitutes a feature (Journal of Neural Engineering, 24 February 2015, Vol. 12:2, 026008).
The "learning" algorithm identifies the mathematical model -- in effect a boundary -- that best separates images from individuals with Parkinson's disease and those without. Applied to an image of unknown status characterized using the same features as in learning, it classifies the image into one of the two groups.
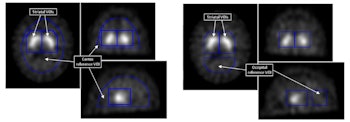
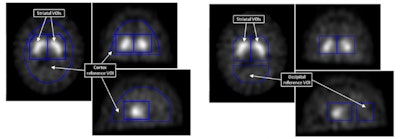
Binding potentials from voxels in striatal VOI were used as features in the support vector machines technique. Cortex and occipital reference VOI were used to calculate them.
Rather than use counts or uptake ratios, the approach analyses binding potentials derived from each striatal voxel, which are a function of the number of dopamine transporters it contains. It equals the counts in that voxel -- that are specifically from tracer molecules bound to dopamine transporters -- normalized to the mean counts per voxel in a reference VOI. The reference VOI contains no dopamine transporters and therefore no transporter-bound tracer. The parameter is a robust measure that varies less than other uptake parameters with factors like the choice of reconstruction algorithm and attenuation correction method.
Large-scale validation
To validate their approach, Castelo-Branco and Oliveira used scans from 209 controls and 445 individuals with early-stage Parkinson's disease. The images were from the Parkinson's Progression Markers Initiative (PPMI), an international, observational, clinical study with the largest dataset of its kind.
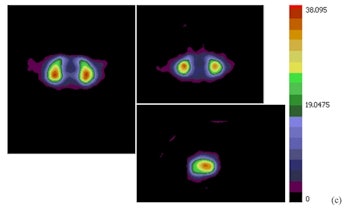
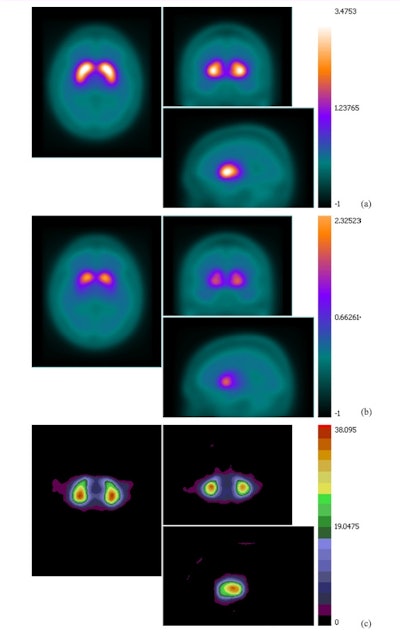
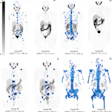
A mean binding potential image for the control group (a) has higher uptake than a mean binding potential image for the Parkinson's disease group (b). A statistically significant difference between the two was demonstrated using a student's t-test (c; t-value image).
Two fixed-size striatal VOI were applied to each image. They were placed at the same anatomical location each time by registering the images to a template, composite image constructed from multiple normal brain images in which the two VOI had been aligned.
The researchers trialed the approach for several permutations of the technique, varying aspects including the registration algorithm, testing both rigid and affine algorithms, and the reference VOI location used to calculate binding potentials. The variations made little difference to the accuracy of the classifier, which ranged from 96.5% to 97.9%.
There was also little change in accuracy when the number of images used to train the learning algorithm was varied, from 327 to 624 images, demonstrating that the technique is robust. Supporting the findings, subsequent, unpublished tests by the researchers using other, smaller, image datasets have produced similar accuracies, as have different reconstruction and attenuation correction methods.
"Based on our results, and also the results from other research groups, we think that it is time for the physicians to give more importance to multivariate analysis-based methods, which can be a powerful complement to the visual analysis or the simple commonly used uptake ratios," Castelo-Branco said.
Both visual and computer-aided assessments of I-123 FP-CIT scans are recommended by the European Association of Nuclear Medicine (EANM) Neuroimaging Committee. An approximate accuracy of 91% has been reported in the literature using visual assessments.
The researchers are developing similar approaches to detect other neurodegenerative conditions including Alzheimer's and Huntingdon's diseases, and plan to apply their technique to PET images using other tracers.
© IOP Publishing Limited. Republished with permission from medicalphysicsweb, a community website covering fundamental research and emerging technologies in medical imaging and radiation therapy.