Top Story
Latest News
First patient dosed with ITM-31 in glioblastoma trial
April 23, 2024
ESR releases movie to mark key moments of ECR 2024
April 23, 2024
Cases of the Week
Check out our Cases of the Week!
More from AuntMinnieEurope
Blackford, Annalise.ai enter partnership
April 23, 2024
RCR: Radiologist shortage continues
April 22, 2024
Accuray opens training facility in Switzerland
April 19, 2024
Raigmore Hospital in Scotland selects RayCare
April 17, 2024
Radiology pays tribute to Dutch pioneer Jacob Valk
April 18, 2024
Parizel makes compelling case for collaboration
April 16, 2024
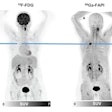
FAPI-PET shows promise in patients with lung cancer
April 16, 2024
Bayer and Hologic collaborate on CEM
April 15, 2024
Radiographers play pivotal research role in Gothenburg
April 15, 2024